Human Breast Cancer IMC 3D Reconstruction#
This tutorial presents an end-to-end DeepSpatial workflow for reconstructing a 3D volume from serial IMC slices of human breast cancer tissue.
In this tutorial, you will:
Import dependencies.
Load and organize slice-level IMC data.
Set up inputs and train DeepSpatial.
Reconstruct the 3D tissue volume.
Visualize and inspect reconstruction results.
1. Import Dependencies#
Import DeepSpatial, utility libraries, and visualization functions used throughout this tutorial.
The tutorial below follows a complete from-scratch workflow.
import os
import re
import glob
import torch
import scanpy as sc
from deepspatial import DeepSpatial
from deepspatial.vis_utils import interactive_3d_labels, interactive_3d_expression, plot_z_distribution, plot_orthogonal_projections
2. Data Preparation#
Please download the Human breast cancer IMC dataset first.
This section scans all h5ad files and uses obsm['spatial_3d'] as the only geometry source.
For each slice, x and y are stored in obsm['spatial'], and z is stored in obs['z_coord'].
# 2. Data preparation and path parsing
# Set this to your local dataset directory before running
data_dir = "/data/yuhangyang/DeepSpatial/data/imc_human_breastcancer"
file_paths = sorted(
glob.glob(os.path.join(data_dir, "imc_*.h5ad")),
key=lambda x: int(re.search(r'imc_(\d+)', os.path.basename(x)).group(1)),
)
if len(file_paths) == 0:
raise FileNotFoundError(f"No files found under: {data_dir}")
# Load slices and split spatial_3d into spatial(x, y) + z_coord
adata_list = []
for p in file_paths:
adata = sc.read_h5ad(p)
if 'spatial_3d' not in adata.obsm or adata.obsm['spatial_3d'].shape[1] < 3:
raise ValueError(
f"{os.path.basename(p)} is missing valid obsm['spatial_3d'] with at least 3 columns"
)
coords_3d = adata.obsm['spatial_3d']
adata.obsm['spatial'] = coords_3d[:, :2].copy()
adata.obs['z_coord'] = coords_3d[:, 2].astype(float)
adata_list.append(adata)
if len(adata_list) == 0:
raise ValueError("No valid IMC slices loaded.")
print(f"Loaded {len(adata_list)} slices with z_coord derived from spatial_3d")
Loaded 15 slices with z_coord derived from spatial_3d
Check Loaded Data#
Inspect loaded slices and confirm spatial alignment before training.
adata_list
[AnnData object with n_obs × n_vars = 7787 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 6931 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 6812 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 6719 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 6739 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 6650 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 6837 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7205 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7181 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7555 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7161 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7301 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7318 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7122 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances',
AnnData object with n_obs × n_vars = 7388 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'X_pca', 'X_umap', 'spatial', 'spatial_3d'
varm: 'PCs'
obsp: 'connectivities', 'distances', 'spatial_connectivities', 'spatial_distances']
3. Setup and Train#
Configure model inputs, build the architecture, and run training in this section.
model = DeepSpatial()
# 3. Data setup: normalization is handled internally without overwriting raw coordinates
candidate_label_keys = ['cell_type', 'cell_class', 'annotation', 'Harmony_labels']
label_key = next((k for k in candidate_label_keys if k in adata_list[0].obs.columns), None)
if label_key is None:
raise ValueError(f'No valid label key found. Tried: {candidate_label_keys}')
print('Using label key:', label_key)
model.setup_data(
adata_list=adata_list,
spatial_key='spatial',
z_key='z_coord',
label_key=label_key,
batch_size=2048
)
Using label key: cell_type
DeepSpatial: Building Trajectories: 100%|██████████| 14/14 [00:36<00:00, 2.62s/it]
Build the Model#
Define architecture and optimizer-related hyperparameters, then train the model. You can adjust settings based on data size, GPU memory, and training goals.
# Build model: configure architecture hyperparameters
model.build_model(
patch_size=8,
hidden_size=256,
depth=6,
lr=2e-4
)
Train the Model#
After training, checkpoints are saved to save_dir. Increase epochs based on convergence in real experiments.
# Train: set checkpoint directory and device options
accelerator = 'gpu' if torch.cuda.is_available() else 'cpu'
devices = [0] if accelerator == 'gpu' else 1
print('Training accelerator:', accelerator)
model.fit(
max_epochs=10,
save_dir="./checkpoints/deepspatial_run",
accelerator=accelerator,
devices=devices,
save_ckpt=False
)
Training accelerator: gpu
GPU available: True (cuda), used: True
TPU available: False, using: 0 TPU cores
💡 Tip: For seamless cloud logging and experiment tracking, try installing [litlogger](https://pypi.org/project/litlogger/) to enable LitLogger, which logs metrics and artifacts automatically to the Lightning Experiments platform.
You are using a CUDA device ('NVIDIA GeForce RTX 4090 D') that has Tensor Cores. To properly utilize them, you should set `torch.set_float32_matmul_precision('medium' | 'high')` which will trade-off precision for performance. For more details, read https://pytorch.org/docs/stable/generated/torch.set_float32_matmul_precision.html#torch.set_float32_matmul_precision
/data/yuhangyang/miniconda3/envs/test-ds/lib/python3.10/site-packages/pytorch_lightning/callbacks/model_checkpoint.py:881: Checkpoint directory /data/yuhangyang/DeepSpatial/docs/source/tutorials/checkpoints/deepspatial_run exists and is not empty.
LOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [0,1,2,3,4,5,6,7]
┏━━━┳━━━━━━━━━━━┳━━━━━━┳━━━━━━━━┳━━━━━━━┳━━━━━━━┓ ┃ ┃ Name ┃ Type ┃ Params ┃ Mode ┃ FLOPs ┃ ┡━━━╇━━━━━━━━━━━╇━━━━━━╇━━━━━━━━╇━━━━━━━╇━━━━━━━┩ │ 0 │ model │ GiT │ 7.7 M │ train │ 0 │ │ 1 │ ema_model │ GiT │ 7.7 M │ eval │ 0 │ └───┴───────────┴──────┴────────┴───────┴───────┘
Trainable params: 7.7 M Non-trainable params: 7.7 M Total params: 15.4 M Total estimated model params size (MB): 61 Modules in train mode: 158 Modules in eval mode: 158 Total FLOPs: 0
/data/yuhangyang/miniconda3/envs/test-ds/lib/python3.10/site-packages/pytorch_lightning/utilities/_pytree.py:21: `isinstance(treespec, LeafSpec)` is deprecated, use `isinstance(treespec, TreeSpec) and treespec.is_leaf()` instead.
/data/yuhangyang/miniconda3/envs/test-ds/lib/python3.10/site-packages/pytorch_lightning/loops/fit_loop.py:534: Found 158 module(s) in eval mode at the start of training. This may lead to unexpected behavior during training. If this is intentional, you can ignore this warning.
`Trainer.fit` stopped: `max_epochs=10` reached.
4. Reconstruct#
Generate a full 3D volume from all IMC slices.
adata_3d = model.reconstruct_full_volume(
adata_list,
thickness=2
)
DeepSpatial: 3D Reconstruct: 100%|██████████| 14/14 [02:01<00:00, 8.67s/gap, range=260.0-280.0um]
Save Reconstruction#
Persist the reconstructed result for future analysis and reproducibility.
os.makedirs("output", exist_ok=True)
adata_3d.write_h5ad("output/deepspatial_3d_imc_breastcancer.h5ad")
print("Saved reconstruction to output/deepspatial_3d_imc_breastcancer.h5ad")
5. Visualization#
Use both interactive and static plots to inspect 3D structure, spatial patterns, and label or gene expression distributions.
adata_3d
AnnData object with n_obs × n_vars = 991185 × 25
obs: 'leiden', 'phenograph', 'cell_type', 'spatial_z', 'z_coord', 'z_norm'
uns: 'leiden', 'leiden_colors', 'moranI', 'neighbors', 'pca', 'spatial_neighbors', 'umap'
obsm: 'spatial'
3D Label Visualization#
Color by the selected label column to quickly verify whether reconstructed cellular patterns are spatially reasonable.
label_candidates = ['cell_type', 'cell_class', 'annotation', 'Harmony_labels']
vis_label = next((k for k in label_candidates if k in adata_3d.obs.columns), None)
if vis_label is None:
raise ValueError(f'No visualization label column found. Tried: {label_candidates}')
interactive_3d_labels(
adata_3d,
color_col=vis_label,
title=f'DeepSpatial IMC Reconstruction ({vis_label})',
width=1000,
height=1000,
)
3D Gene Expression Visualization#
Map a representative marker gene into 3D space to identify local enrichment patterns.
gene_name = adata_3d.var_names[0]
for g in ['PanCK', 'CD45', 'Ki67']:
if g in adata_3d.var_names:
gene_name = g
break
interactive_3d_expression(
adata_3d,
gene_name=gene_name,
title=f'IMC Reconstruction Expression: {gene_name}',
width=1000,
height=1000,
)
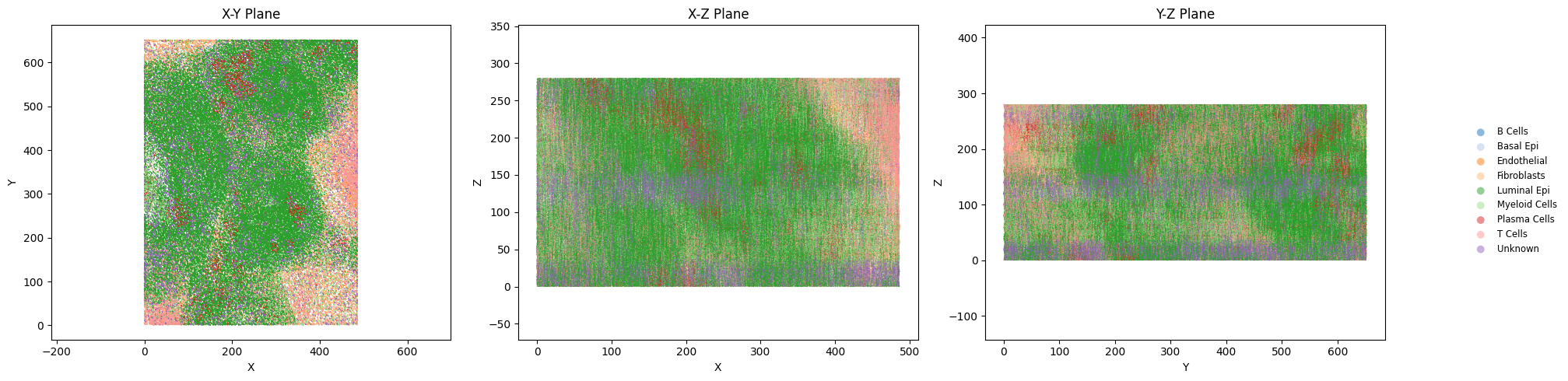
Optional: Orthogonal and Z-Distribution Checks#
Use additional plots to inspect spatial layout consistency along XY, XZ, YZ and Z-axis distributions.
label_candidates = ['cell_type', 'cell_class', 'annotation', 'Harmony_labels']
vis_label = next((k for k in label_candidates if k in adata_3d.obs.columns), None)
plot_orthogonal_projections(adata_3d, color_col=vis_label)
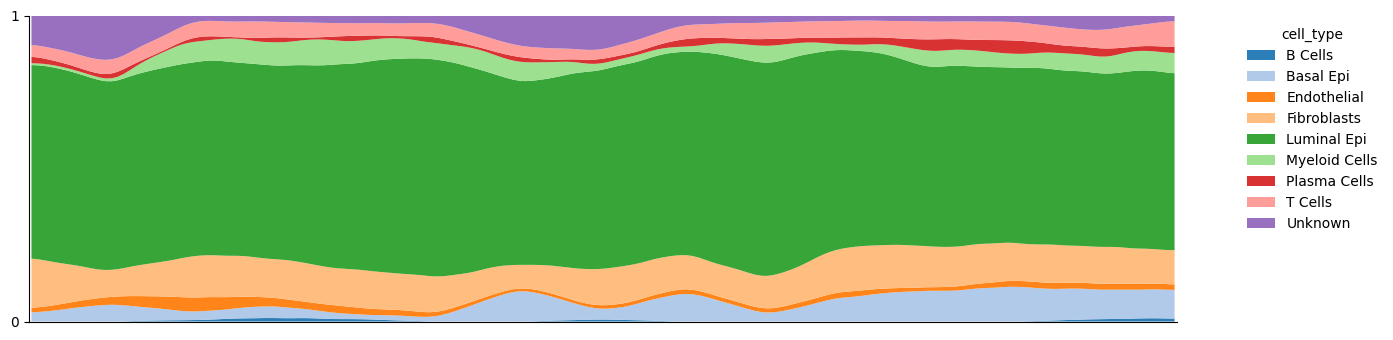
plot_z_distribution(adata_3d, color_col=vis_label, smooth_sigma=2)