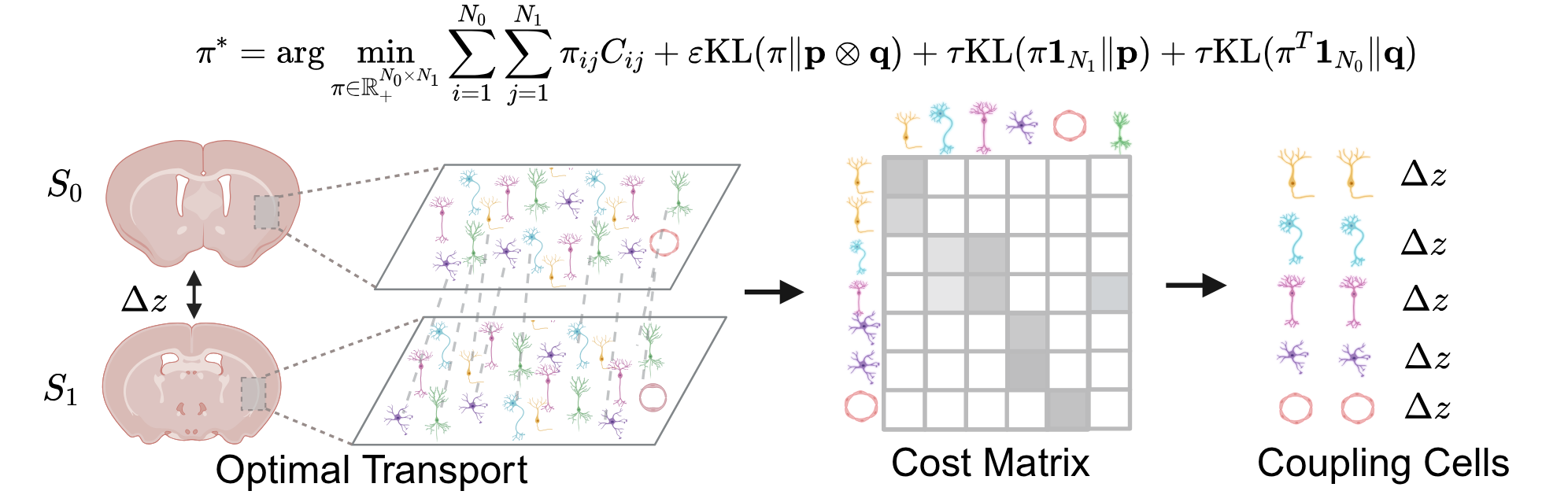
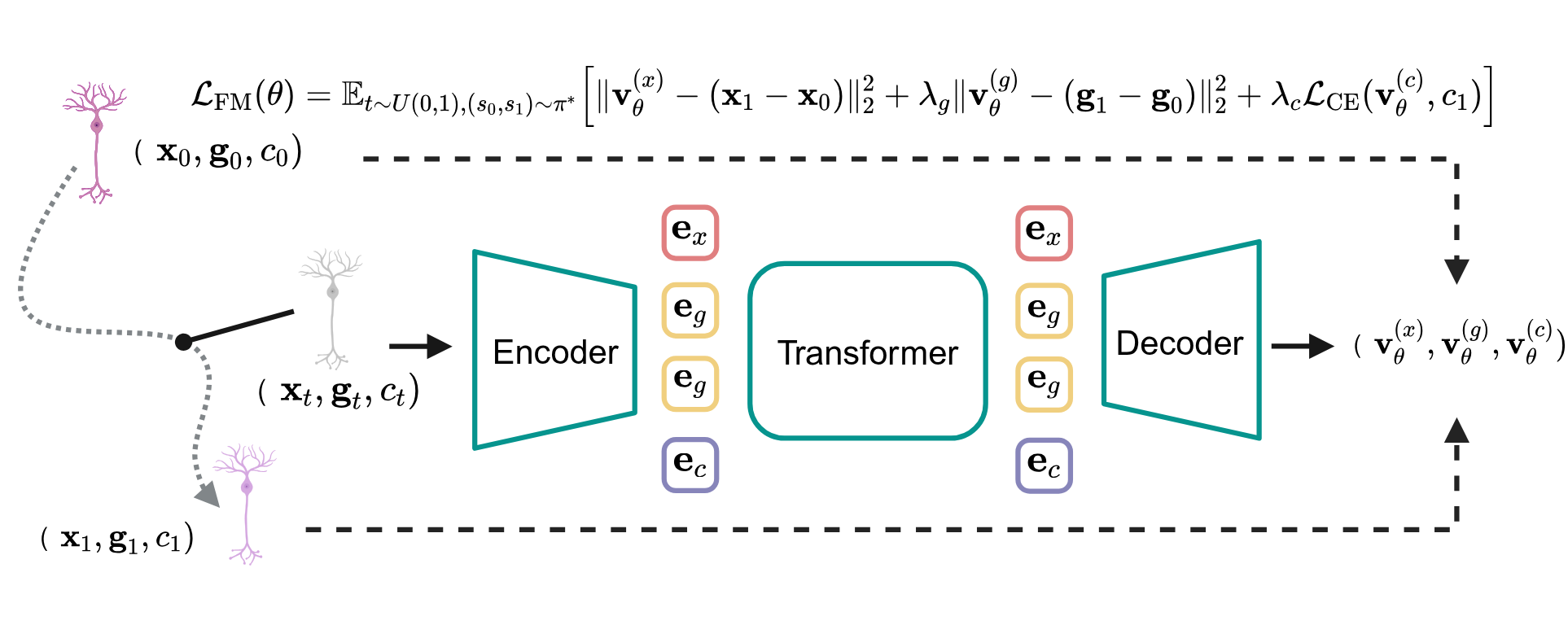
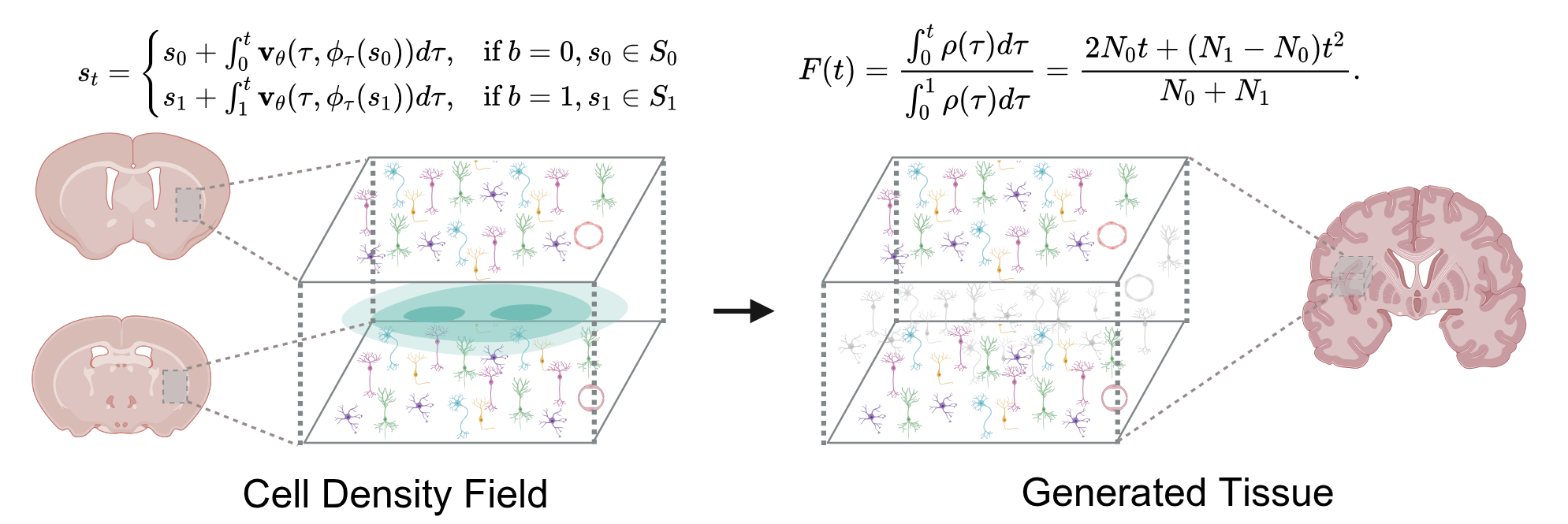
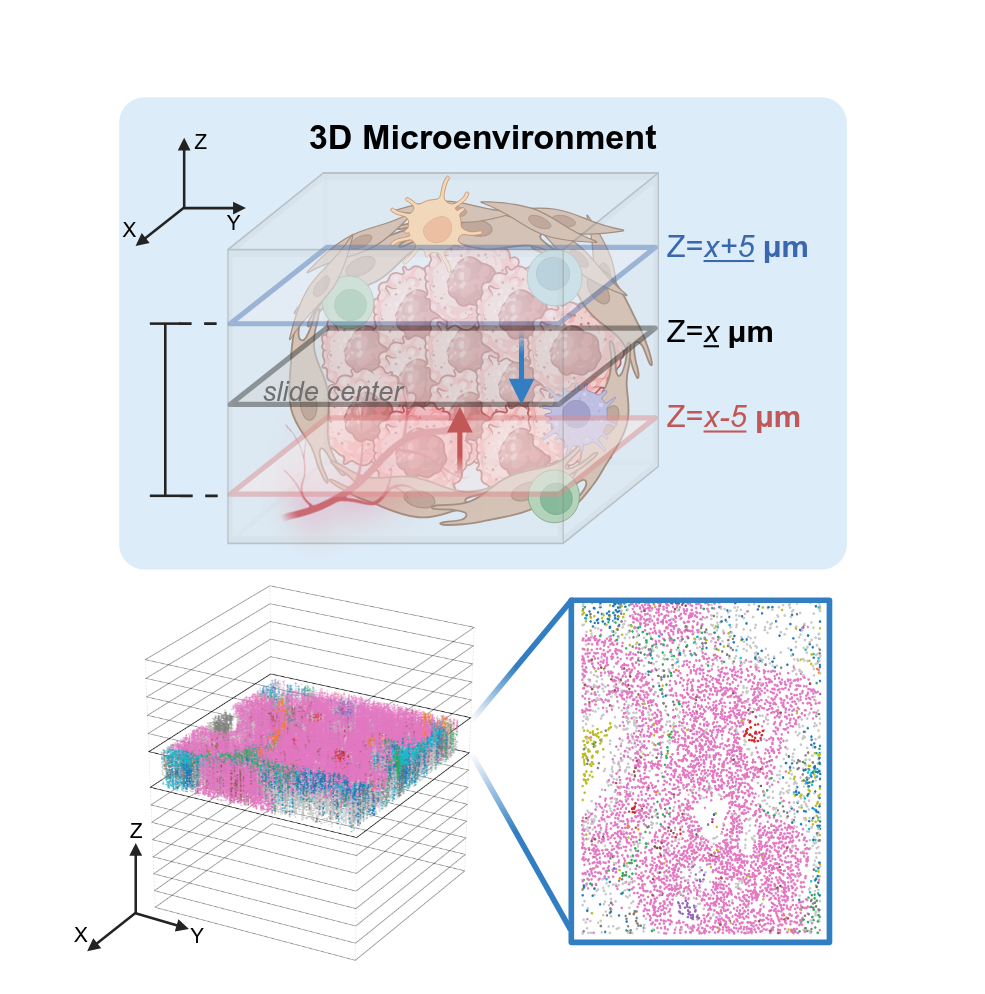
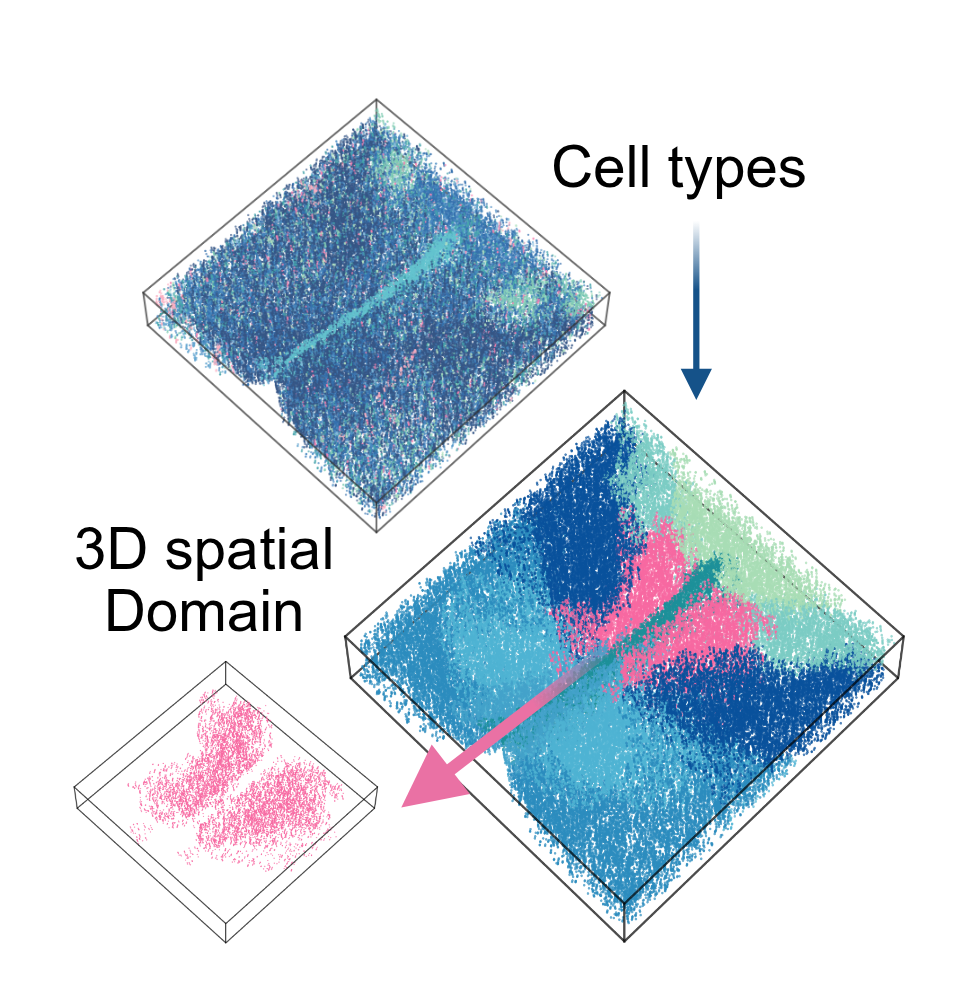
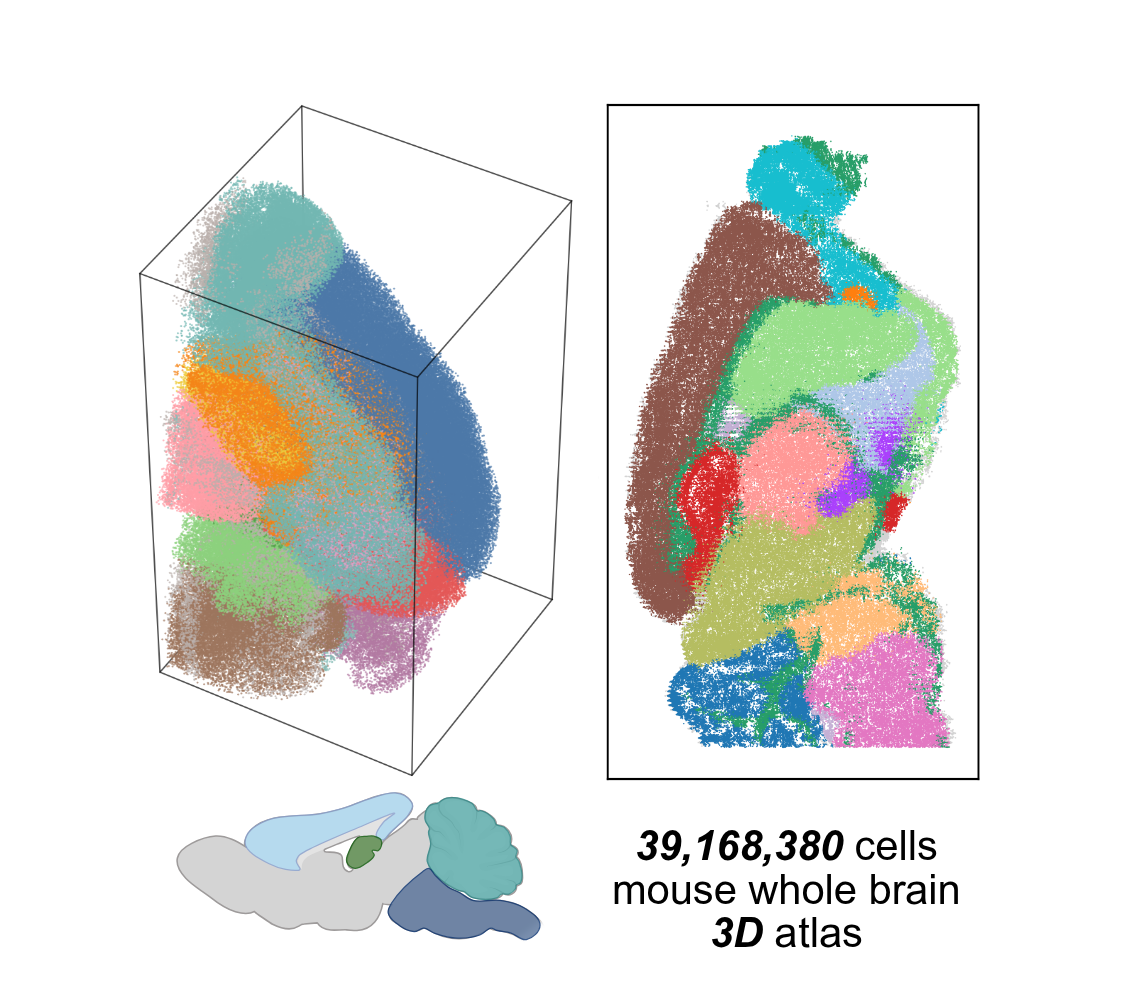
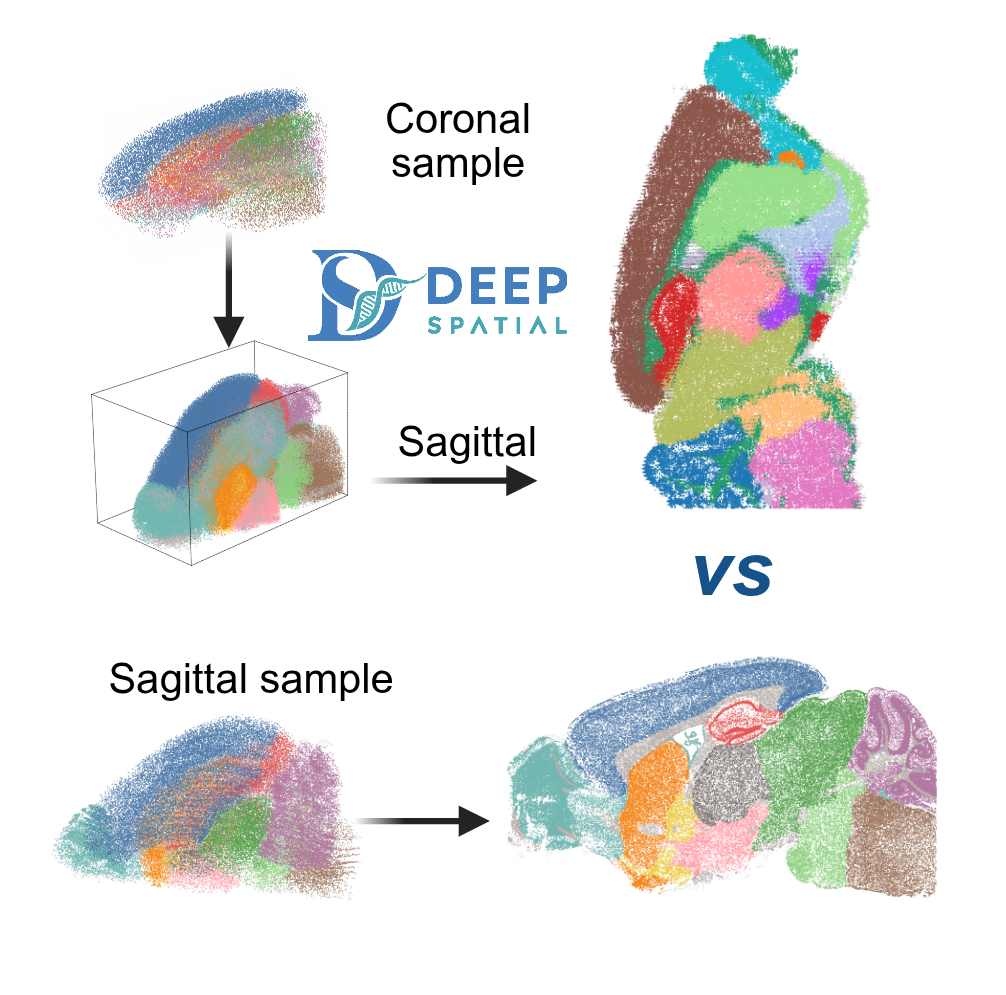
From Discrete Slices to Continuous Volume
DeepSpatial reframes 3D reconstruction as a continuous probability density evolution. Rather than merely inserting pseudo-2D planes, it models the multi-modal transitions between physical slices as mixed continuous-discrete dynamics via a scalable Transformer architecture.